- Overview
- Download
- Updates
- License
- Tutorial
- Case Tests
- Contacts
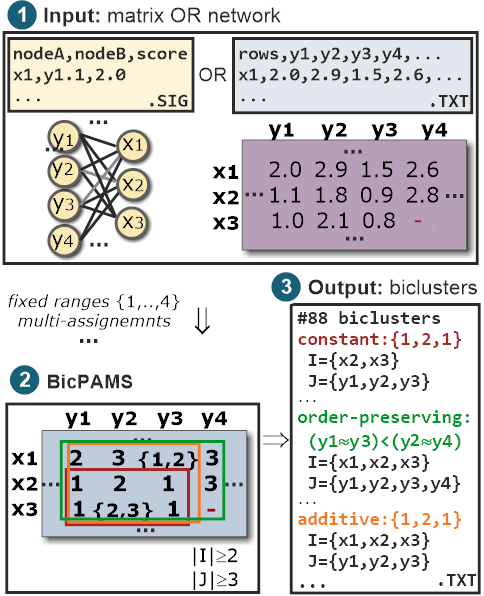
BicPAMS is a tool for the exploratory and unsupervised analysis of tabular and network data.
BicPAMS integrates state-of-the-art contributions on pattern-based biclustering (including BicPAM, BicNET, BicSPAM, BiC2PAM, BiP, DeBi and BiModule).
As such, BicPAMS is able to efficiently discover exhaustive and flexible structures of biclusters, with parameterizable coherency and robustness to noise and missings.
BicPAMS has been successfully tested on biomedical data (including expression data, clinical data and biological networks) and social data.
|
Rui Henriques, Francisco L. Ferreira and Sara C. Madeira, 2017, BicPAMS: Software for Biological Data Analysis with Pattern-based Biclustering, BMC Bioinformatics, 18:82 [10.1186/s12859-017-149] |
|
Rui Henriques and Sara C. Madeira, 2016, BicNET: Flexible module discovery in large-scale biological networks using biclustering, Algorithms for Molecular Biology, 11(1):1--30 [10.1186/s13015-016-0074-8] |
|
Rui Henriques, Claudia Antunes and Sara C. Madeira, 2015, A Structured View on Pattern Mining-based Biclustering,
Pattern Recognition, 48(12):3941-3958, Elsevier [http://dx.doi.org/10.1016/j.patcog.2015.06.018] |
|
Rui Henriques and Sara C. Madeira, 2015, BiC2PAM: constraint-guided biclustering for biological data analysis with domain knowledge,
Algorithms for Molecular Biology 2016, 11:23 [10.1007/978-3-319-23485-4_34] |
|
Rui Henriques and Sara C. Madeira, 2015, Biclustering with Flexible Plaid Models to Unravel Interactions between Biological Processes,
IEEE/ACM Transactions on Computational Biology and Bioinformatics, 12(4):738-752, IEEE/ACM [10.1109/TCBB.2014.2388206] |
|
Rui Henriques and Sara C. Madeira, 2014, BicSPAM: Flexible Biclustering using Sequential Patterns,
BMC Bioinformatics, 15:130, BioMed Central Ltd [doi:10.1186/1471-2105-15-130] |
| Rui Henriques and Sara C. Madeira, 2014, BicPAM: Pattern-based biclustering for biomedical data analysis, Algorithms for Molecular Biology, 9(1):27-, BioMed Central Ltd [10.1186/s13015-014-0027-z] |
| Rui Henriques, Sara C. Madeira and Claudia Antunes, 2013, F2G: Efficient Discovery of Full-Patterns,
In ECML/PKDD IW on New Frontiers to Mine Complex Patterns, Springer-Verlag, Prague, Czech Republic |
| Rui Henriques, Claudia Antunes and Sara C. Madeira, 2014, Methods for the Efficient Discovery of Large Item-Indexable Sequential Patterns,
Lecture Notes in Computer Science, 100-116, Springer International Publishing [10.1007/978-3-319-08407-7_7] |
Desktop GUI : download BicPAMS GUI v4.0.4 (requirement: Java v7 or superior)
Java API: download BicPAMS API v4.0.2 (requirement: JDK v1.7 or superior)
For previous versions of the BicPAMS software please contact the authors.
2017/01/11: The software article describing the BicPAMS software was published in BMC Bioinformatics.
2017/01/11: Methods to assess the statistical significance of biclusters (BSig) were added to the source code (soon available in BicPAMS GUI).
2016/12/08: A stable version of the software (GUI, API, console and source code) was uploaded.
2015/11/21: Desktop graphical interface of BicPAMS was released.
2015/10/27: A stable version of the programmatic interface of BicPAMS was released. This interface consistently combines the behavior of BicNET, BicPAM, BicSPAM, BiP and BiC2PAM.
2014/09/17: A stable version of BicNET was released for biclustering network data in order to retrieve non-trivial yet meaningful modules.
2014/01/21: A stable version of BiC2PAM was released for constraint-based biclustering in the presence of background knowledge.
2013/07/20: A stable version of BiP was released for biclustering data with plaid effects.
2012/06/13: A stable version of BicSPAM was released for flexible and robust order-preserving biclustering.
2012/04/21: A stable version of BicPAM was released for flexible and robust biclustering with constant, additive, multiplicative and symmetric coherency assumptions.
BicPAMS software is made solely available for research and educational purposes.
In this context, you can use the software under: 1) the terms of the GNU General Public License as published by the Free Software Foundation (either version 3 or any later version), and 2) its proper citation.
BicPAMS is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY (without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE).
If you wish to use it for other purposes, you must contact the authors for permission.
Introductory pointers
Illustrative Example