This exemplifies the matlab fuctions used to calculate the entropic profiles
fastafile='m4.seq';
fasta=readfasta(fastafile);
%concatenate sequences seq=[]; for i=1:length(fasta.sequence); seq=[seq,fasta.sequence{i}]; end L=length(seq)
L =
2000
[cf,cb]=count_repeat(seq);
Forward counts... Backward counts...
[KM,Ns,phis]=fill_kernel3D(cf); %use default parameters (forward counts) %study a particular N=6 for all phis figure; plot(squeeze(KM(6,:,:))','.-'); xlabel('position'); ylabel('kernel density'); title([fasta.title ' N=6; \phi variable']); legend(num2str((1:10)'),2); % %study KM=f(pos,Ns) for fixed phis % for i=1:12; % figure % plot(squeeze(KM(:,i,:))','.-'); % xlabel('position'); % ylabel('kernel density'); % title([fasta.title ' \phi=' num2str(i) '; N variable']); % legend(num2str((1:10)'),2); % end %study KM=f(pos,Ns) for fixed phis figure plot(squeeze(KM(:,12,:))','.-'); xlabel('position'); ylabel('kernel density'); title([fasta.title ' \phi=' num2str(12) '; N variable']); legend(num2str((1:10)'),2);

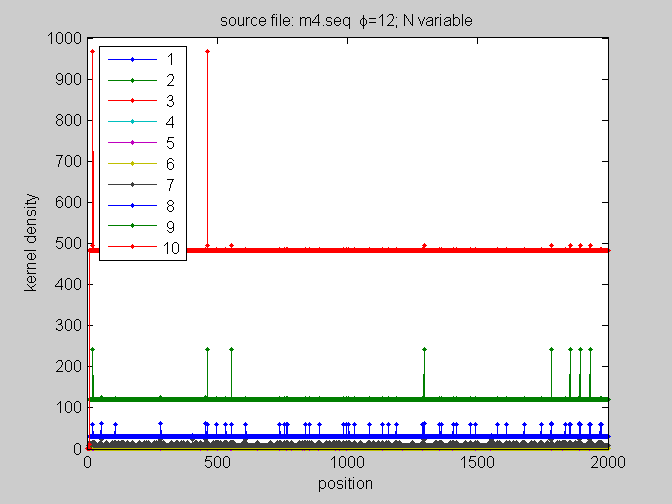
KMn=normKM(KM,1);
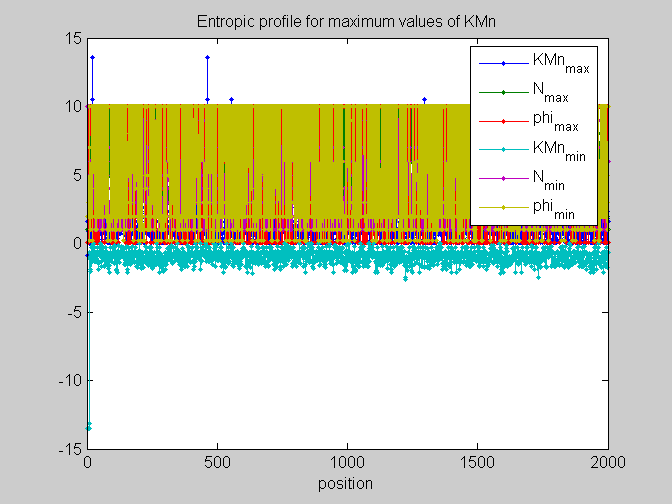
[pos,code]=find_scale(KMn,Ns,phis); %...and plot figure;plot(pos,'.-');legend(code);xlabel('position');ylabel('kernel density') %partial plots figure;plot(pos(:,[1,4]),'.-');legend(code([1,4]));xlabel('position');ylabel('kernel density') figure;plot(pos(:,[2,5]),'.-');legend(code([2,5]));xlabel('position');ylabel('kernel density') figure;plot(pos(:,[3,6]),'.-');legend(code([3,6]));xlabel('position');ylabel('kernel density')
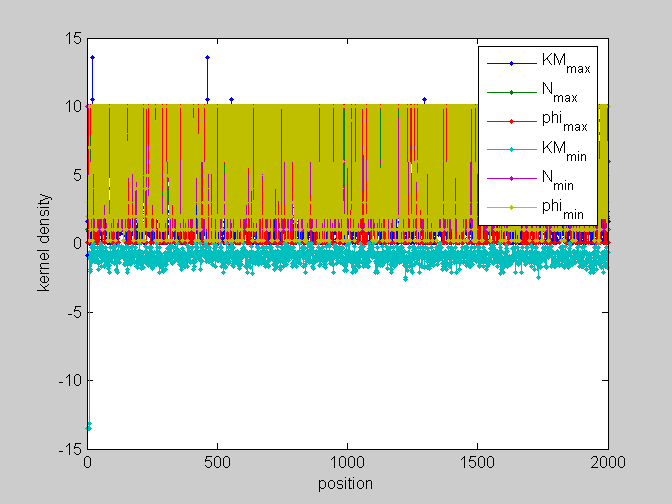
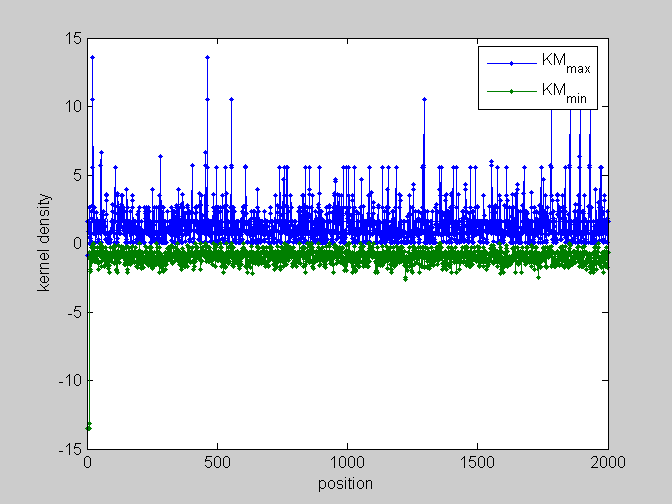
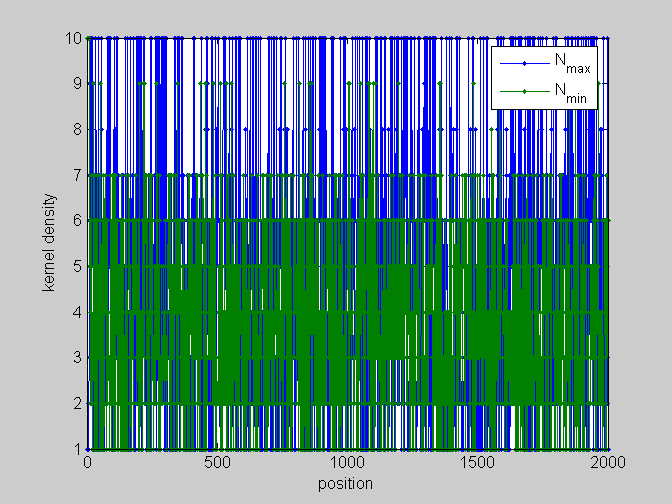
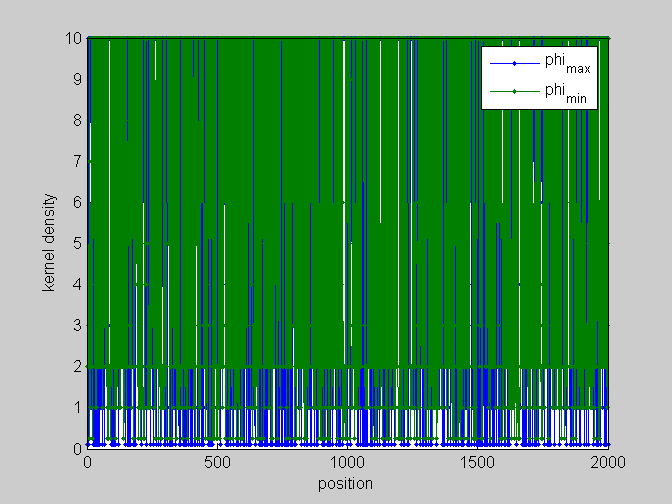
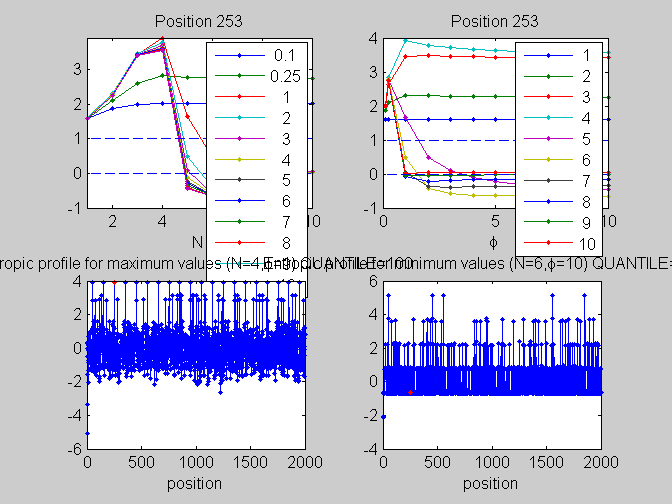
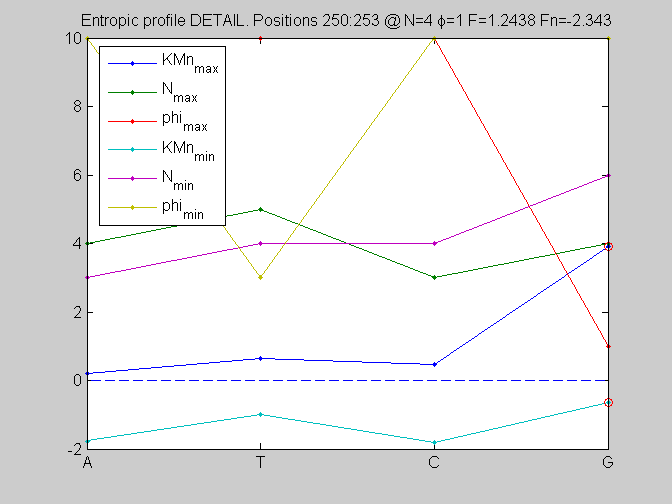
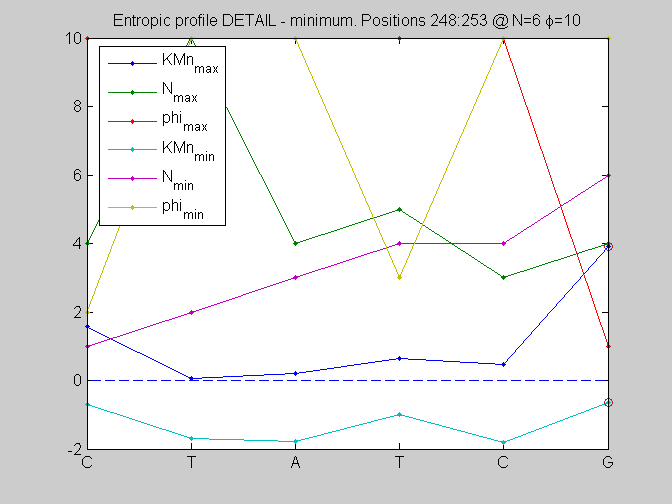
local_study(253,KMn,phis,Ns,pos,L,seq,KM);
F =
1.2438
Fn =
-2.3430